GenBank/NCBI API MCP Server for Claude Code 4 tools — connect in under 2 minutes
Claude Code is Anthropic's agentic CLI for terminal-first development. Add GenBank/NCBI API as an MCP server in one command and Claude Code will discover every tool at runtime. ideal for automation pipelines, CI/CD integration, and headless workflows via Vinkius.
ASK AI ABOUT THIS MCP SERVER
Vinkius supports streamable HTTP and SSE.
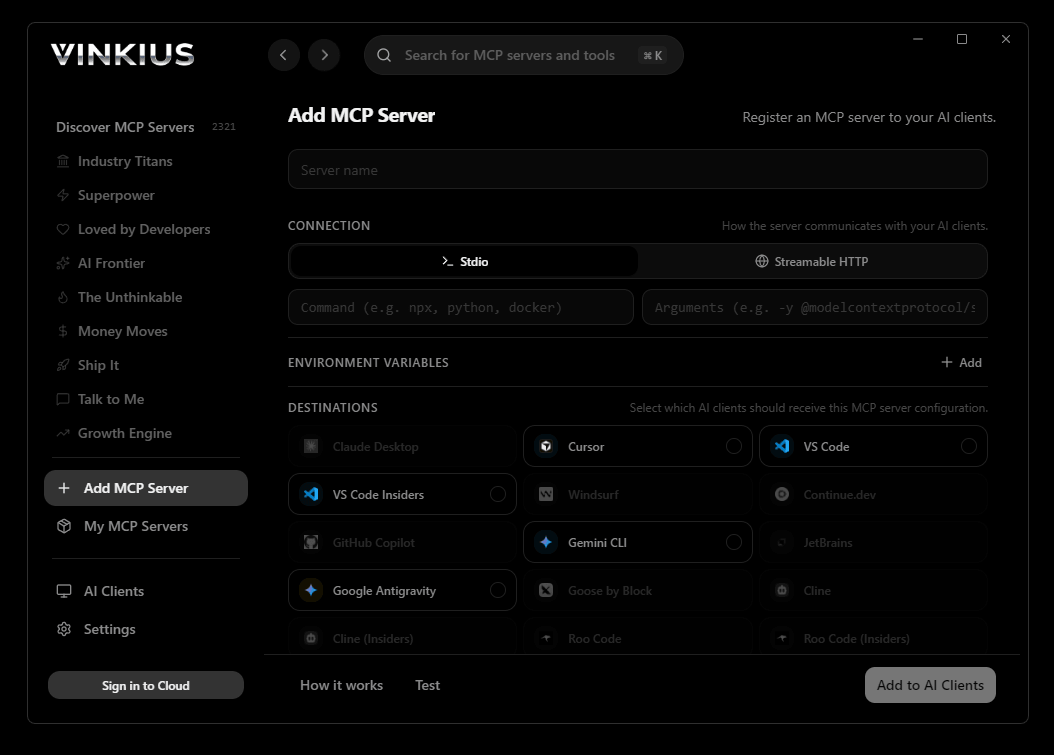
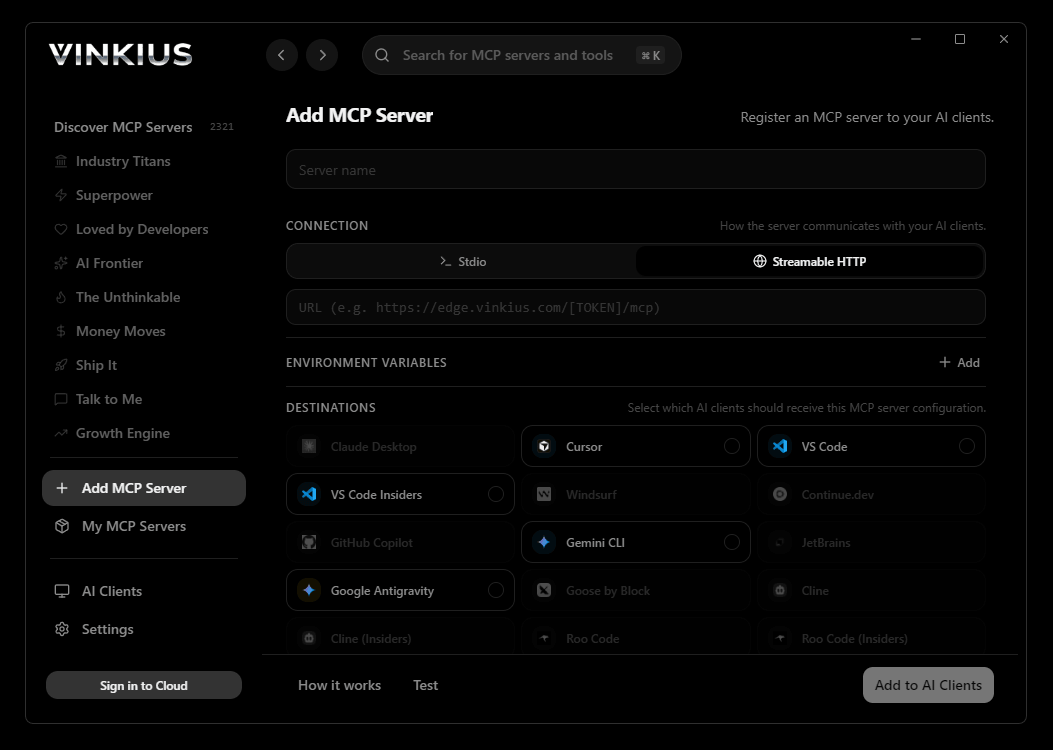
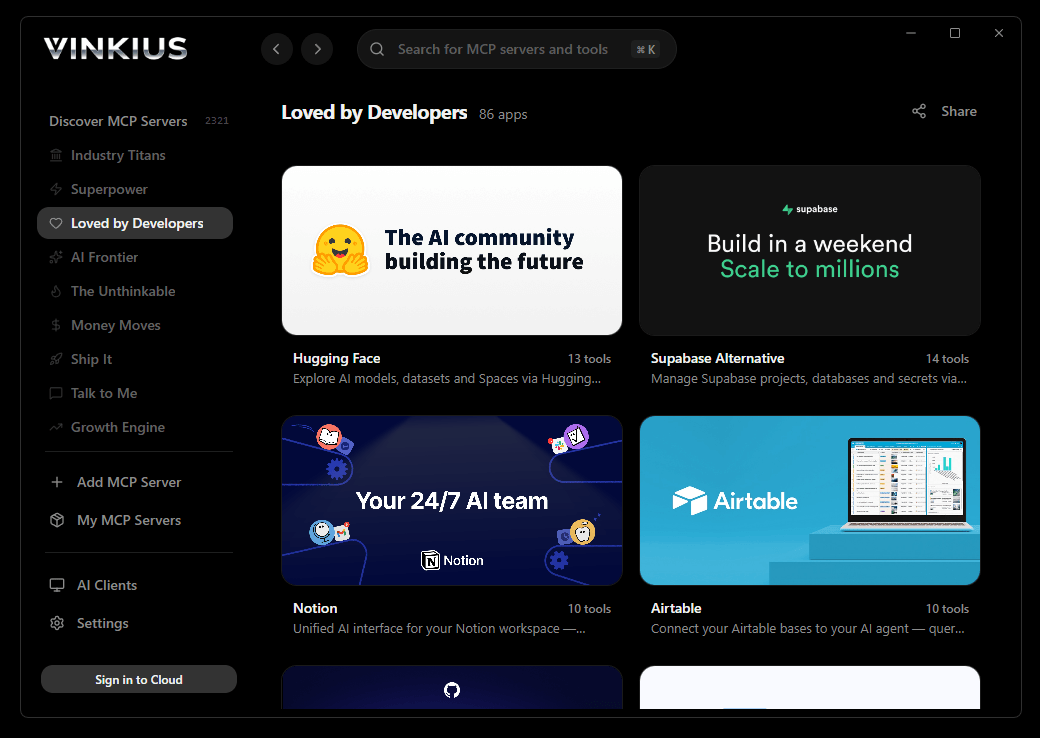
Vinkius Desktop App
The modern way to manage MCP Servers — no config files, no terminal commands. Install GenBank/NCBI API and 2,500+ MCP Servers from a single visual interface.

# Your Vinkius token. get it at cloud.vinkius.com
claude mcp add genbankncbi-api --transport http "https://edge.vinkius.com/[YOUR_TOKEN_HERE]/mcp"
* Every MCP server runs on Vinkius-managed infrastructure inside AWS - a purpose-built runtime with per-request V8 isolates, Ed25519 signed audit chains, and sub-40ms cold starts optimized for native MCP execution. See our infrastructure
About GenBank/NCBI API MCP Server
Empower your AI agent to orchestrate your entire biological research and genomic auditing workflow with the GenBank/NCBI API, the authoritative source for molecular biology data from the National Center for Biotechnology Information. By connecting NCBI's E-utilities to your agent, you transform complex sequence searches into a natural conversation. Your agent can instantly retrieve sequence UIDs, audit bibliographic summaries, and query protein metadata without you ever touching a bioinformatics portal. Whether you are conducting evolutionary research or managing laboratory constraints, your agent acts as a real-time genomic consultant, ensuring your data is always verified and precise.
Claude Code registers GenBank/NCBI API as an MCP server in a single terminal command. Once connected, Claude Code discovers all 4 tools at runtime and can call them headlessly. ideal for CI/CD pipelines, cron jobs, and automated workflows where GenBank/NCBI API data drives decisions without human intervention.
What you can do
- Sequence Auditing — Search for thousands of genetic sequences by keyword and retrieve high-resolution metadata, including UIDs and titles.
- Database Oversight — Audit multiple NCBI databases like 'nuccore' or 'protein' to understand the thematic distribution of biological data instantly.
- Summary Discovery — Query technical and bibliographic summaries for specific sequence IDs to assist in deep-dive archival classification.
- Metadata Intelligence — Retrieve unique identifiers and caption details for any biological record to assist in scientific auditing.
- Operational Monitoring — Check API status to ensure your bioinformatics research workflow is always operational.
The GenBank/NCBI API MCP Server exposes 4 tools through the Vinkius. Connect it to Claude Code in under two minutes — no API keys to rotate, no infrastructure to provision, no vendor lock-in. Your configuration, your data, your control.
How to Connect GenBank/NCBI API to Claude Code via MCP
Follow these steps to integrate the GenBank/NCBI API MCP Server with Claude Code.
Install Claude Code
Run npm install -g @anthropic-ai/claude-code if not already installed
Add the MCP Server
Run the command above in your terminal
Verify the connection
Run claude mcp to list connected servers, or type /mcp inside a session
Start using GenBank/NCBI API
Ask Claude: "Using GenBank/NCBI API, show me...". 4 tools are ready
Why Use Claude Code with the GenBank/NCBI API MCP Server
Claude Code provides unique advantages when paired with GenBank/NCBI API through the Model Context Protocol.
Single-command setup: `claude mcp add` registers the server instantly. no config files to edit or applications to restart
Terminal-native workflow means MCP tools integrate seamlessly into shell scripts, CI/CD pipelines, and automated DevOps tasks
Claude Code runs headlessly, enabling unattended batch processing using GenBank/NCBI API tools in cron jobs or deployment scripts
Built by the same team that created the MCP protocol, ensuring first-class compatibility and the fastest adoption of new protocol features
GenBank/NCBI API + Claude Code Use Cases
Practical scenarios where Claude Code combined with the GenBank/NCBI API MCP Server delivers measurable value.
CI/CD integration: embed GenBank/NCBI API tool calls in your deployment pipeline to validate configurations or fetch secrets before shipping
Headless batch processing: schedule Claude Code to query GenBank/NCBI API nightly and generate reports without human intervention
Shell scripting: pipe GenBank/NCBI API outputs into other CLI tools for data transformation, filtering, and aggregation
Infrastructure monitoring: run Claude Code in a cron job to query GenBank/NCBI API status endpoints and alert on anomalies
GenBank/NCBI API MCP Tools for Claude Code (4)
These 4 tools become available when you connect GenBank/NCBI API to Claude Code via MCP:
check_api_status
Check if the NCBI E-utilities service is operational
get_ncbi_summary
Get a summary for a specific NCBI sequence ID
list_ncbi_databases
List all available NCBI databases
search_ncbi_sequences
g., nuccore, protein) based on a query term. Search for biological sequences in an NCBI database
Example Prompts for GenBank/NCBI API in Claude Code
Ready-to-use prompts you can give your Claude Code agent to start working with GenBank/NCBI API immediately.
"Search for 'human insulin' in the 'protein' database using NCBI."
"Get the summary for NCBI UID '123456' in 'nuccore'."
"List all available NCBI databases."
Troubleshooting GenBank/NCBI API MCP Server with Claude Code
Common issues when connecting GenBank/NCBI API to Claude Code through the Vinkius, and how to resolve them.
Command not found: claude
npm install -g @anthropic-ai/claude-codeConnection timeout
GenBank/NCBI API + Claude Code FAQ
Common questions about integrating GenBank/NCBI API MCP Server with Claude Code.
How do I add an MCP server to Claude Code?
claude mcp add --transport http "" in your terminal. Claude Code registers the server and discovers all tools immediately.Can Claude Code run MCP tools in headless mode?
How do I list all connected MCP servers?
claude mcp in your terminal to see all registered servers and their status, or type /mcp inside an active Claude Code session.Connect GenBank/NCBI API with your favorite client
Step-by-step setup guides for every MCP-compatible client and framework:
Anthropic's native desktop app for Claude with built-in MCP support.
AI-first code editor with integrated LLM-powered coding assistance.
GitHub Copilot in VS Code with Agent mode and MCP support.
Purpose-built IDE for agentic AI coding workflows.
Autonomous AI coding agent that runs inside VS Code.
Anthropic's agentic CLI for terminal-first development.
Python SDK for building production-grade OpenAI agent workflows.
Google's framework for building production AI agents.
Type-safe agent development for Python with first-class MCP support.
TypeScript toolkit for building AI-powered web applications.
TypeScript-native agent framework for modern web stacks.
Python framework for orchestrating collaborative AI agent crews.
Leading Python framework for composable LLM applications.
Data-aware AI agent framework for structured and unstructured sources.
Microsoft's framework for multi-agent collaborative conversations.
Connect GenBank/NCBI API to Claude Code
Get your token, paste the configuration, and start using 4 tools in under 2 minutes. No API key management needed.
